
Imageomics
Imageomics Future and Emerging Technologies (FET-OPEN) 2020: High-throughput imaging of proteins using nanobodies Horizon 2020 01.07.2021-31.12.2024 The conventional detection of proteins by imaging employs fluorophores
Since 2001, Prof. Dr. Rizzoli focused his work on mechanisms of pre-synaptic function, especially on synaptic vesicle recycling. Over the years, we have identified and developed a new concept of neurotransmitter release, in which a small number of “perfect” synaptic vesicles participate in most synaptic release events. These vesicles are mobile and therefore are always able to move towards the release sites (active zones), while all other vesicles are immobile, and fuse only rarely. We have described various elements of this model (Rizzoli and Betz, Science, 2004; Westphal et al., Science, 2008; Kamin et al., Biophys J, 2010; Hoopmann et al., PNAS, 2010; Wilhelm et al., Nat Neurosci, 2010), and also extended it to most known synaptic preparations in important reviews of the field (Rizzoli and Betz, Nat Rev Neurosci, 2005; updated in Denker and Rizzoli, Front Synaptic Neurosci, 2010). Recently, we demonstrated this concept in synapses of behaving animals, in what is the first study of vesicle function in vivo (Denker et al., PNAS, 2011a; Denker et al., PNAS, 2011b). In addition to the synaptic vesicle studies, we have applied the membrane trafficking tools we developed to other systems such as endosomal sorting (for example Bethani et al., EMBO J, 2007; Barysch et al., PNAS, 2009).

In a second perspective, we have a strong interest in developing microscopy techniques, most importantly by applying super-resolution microscopy to biological questions. Focusing on stimulated emission depletion (STED) microscopy, we provided the first true biological application of the technique (Willig et al., Nature, 2006) and the first live-imaging application (Westphal et al., Science, 2008), besides participating in other efforts such as the first multi-color super-resolution applications (Donnert et al., Biophys J, 2007).
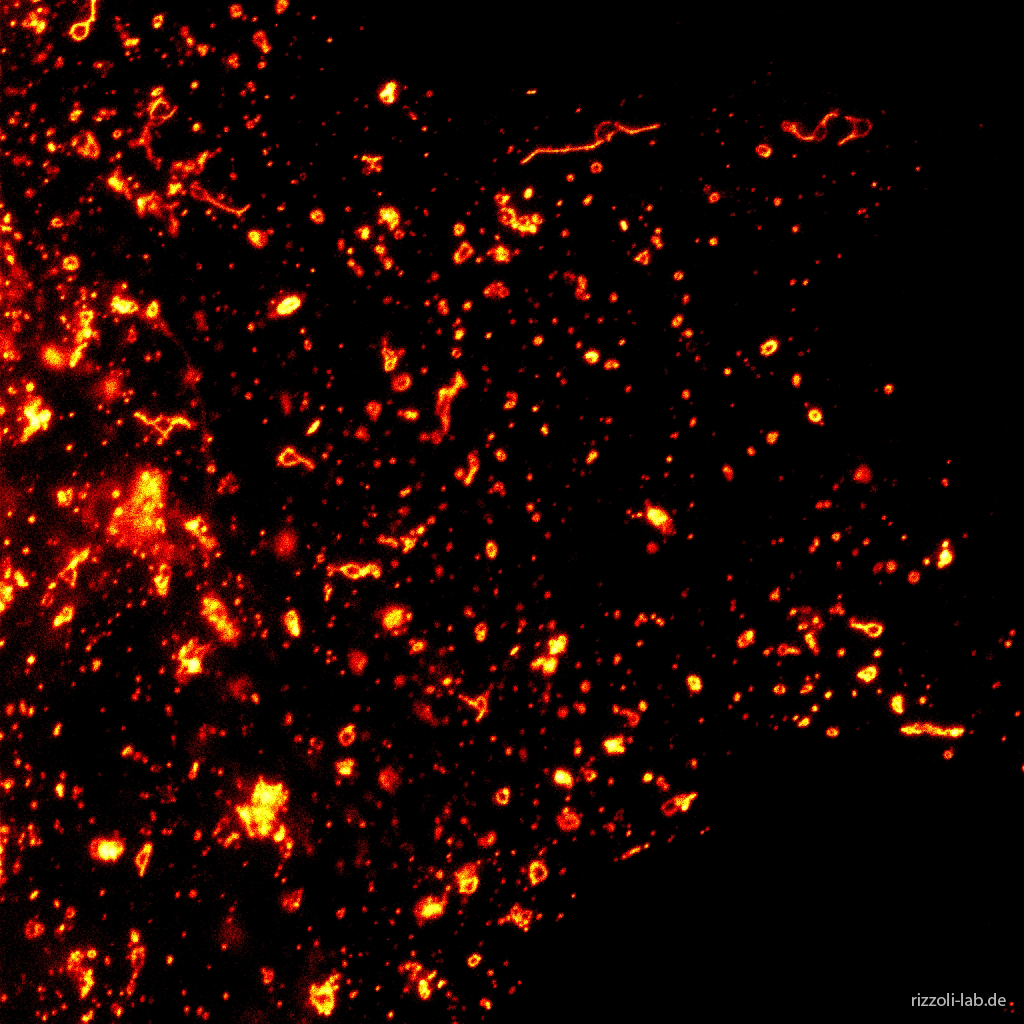

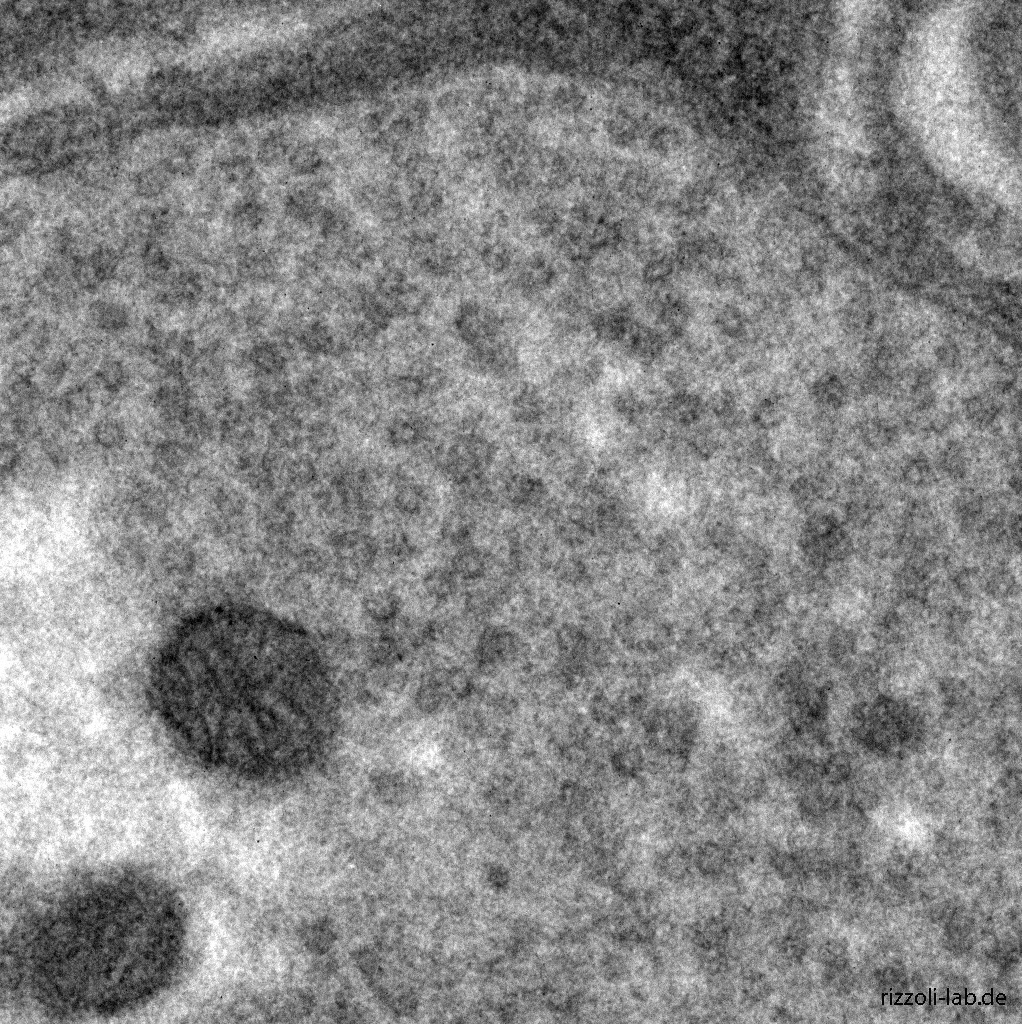
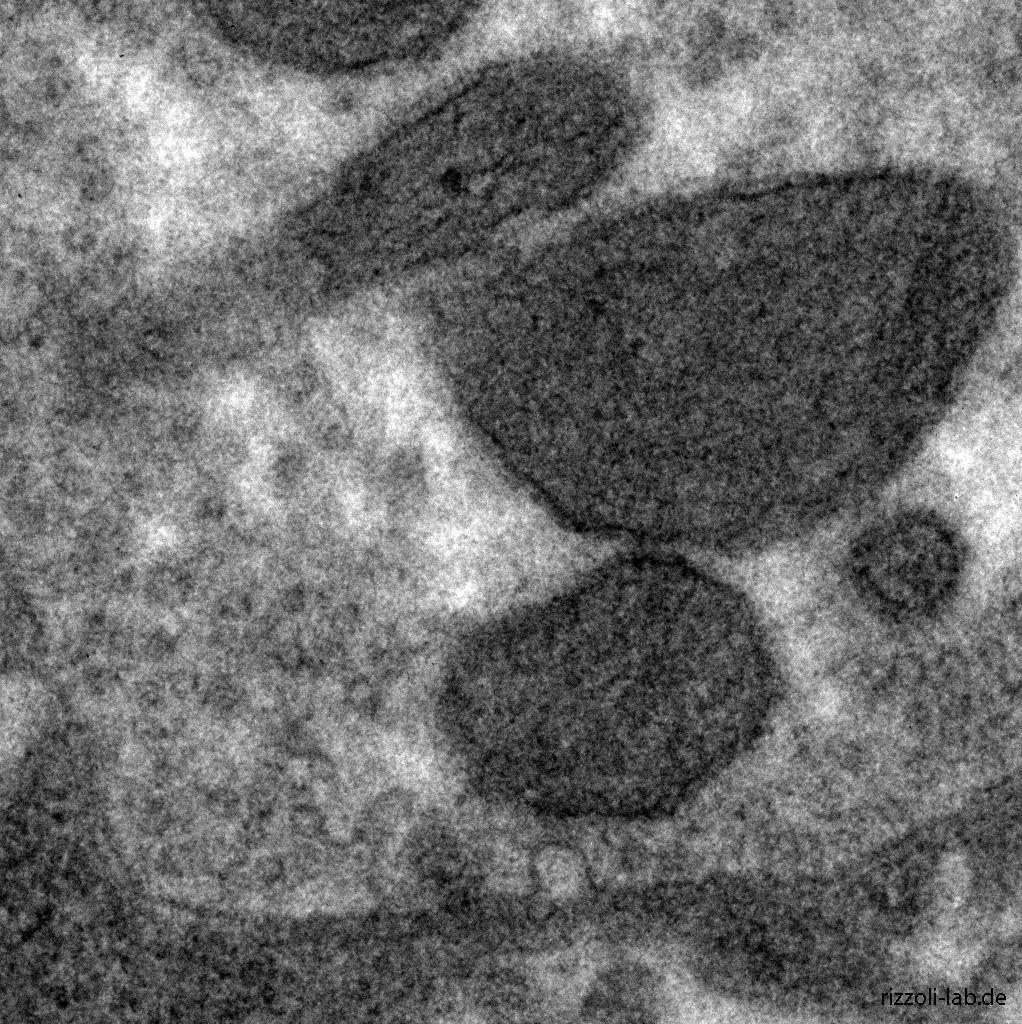
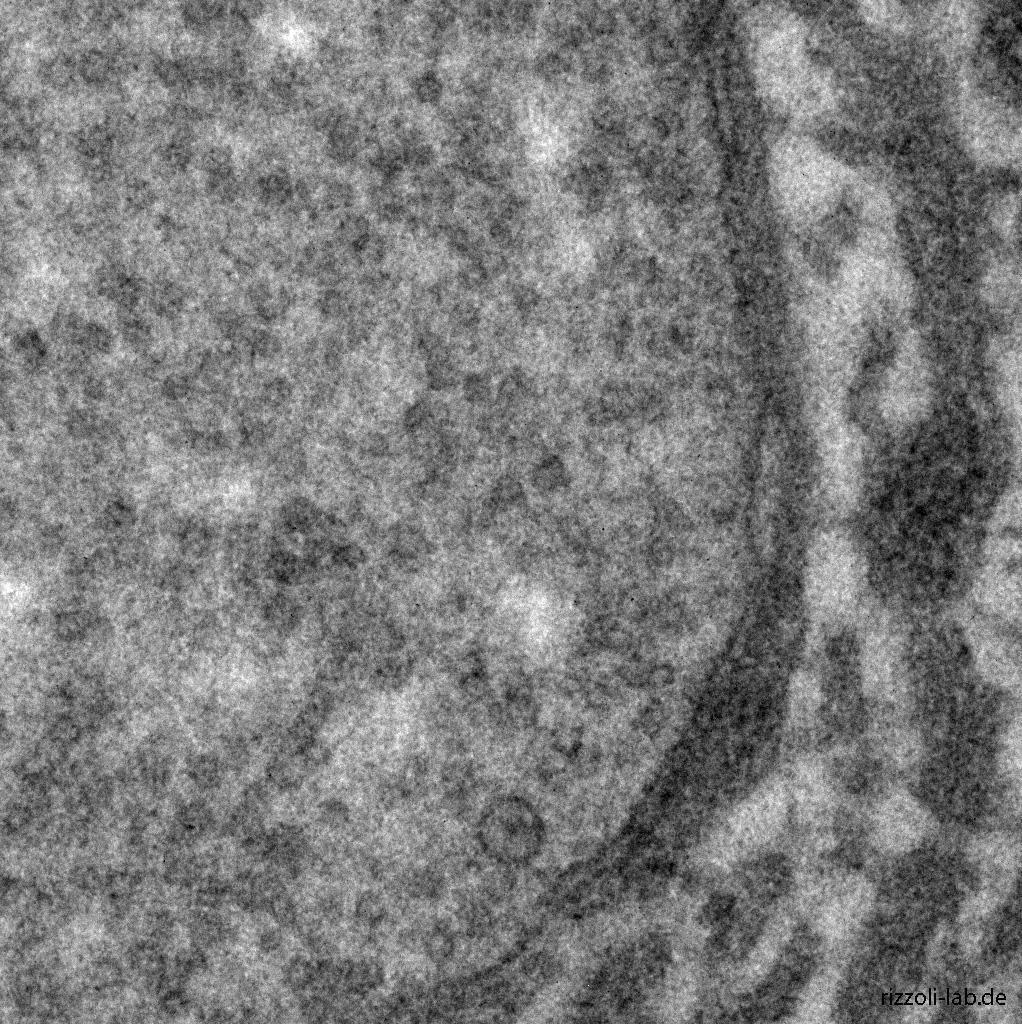
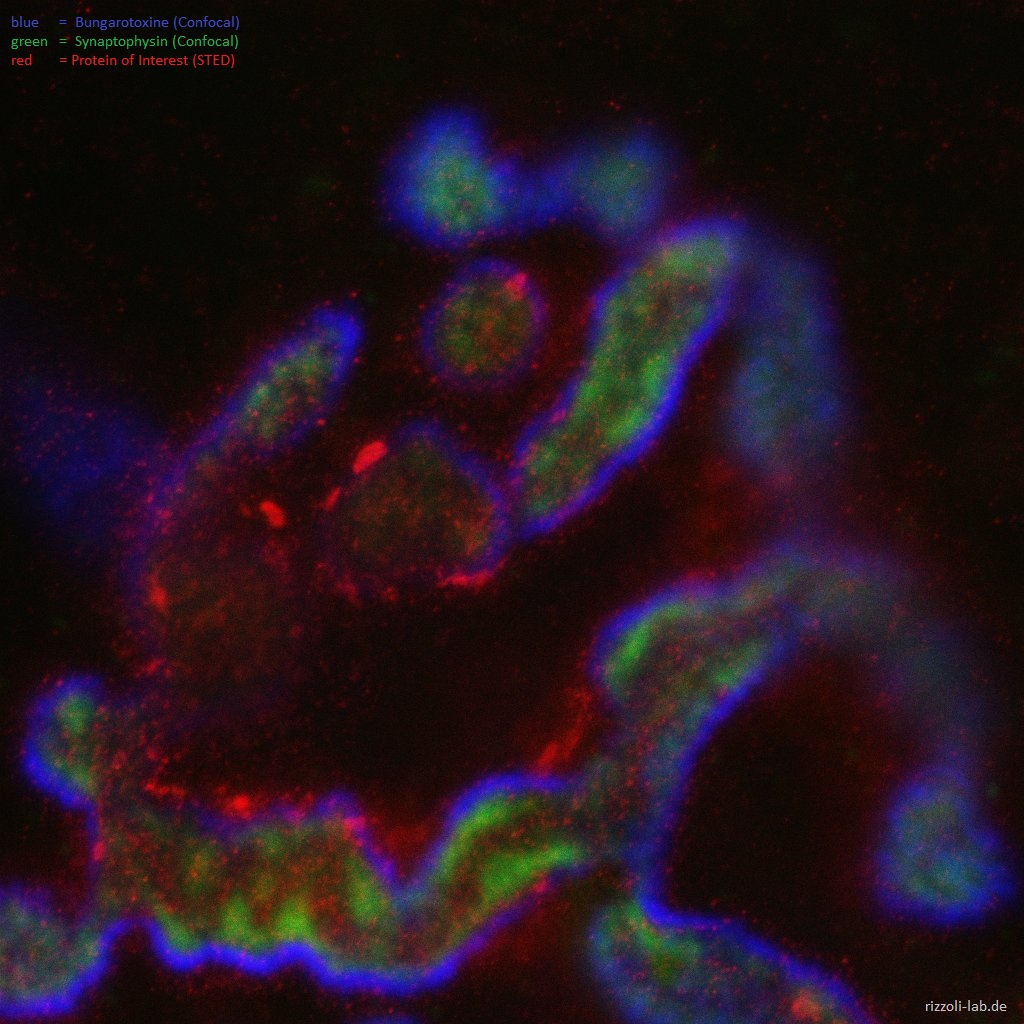
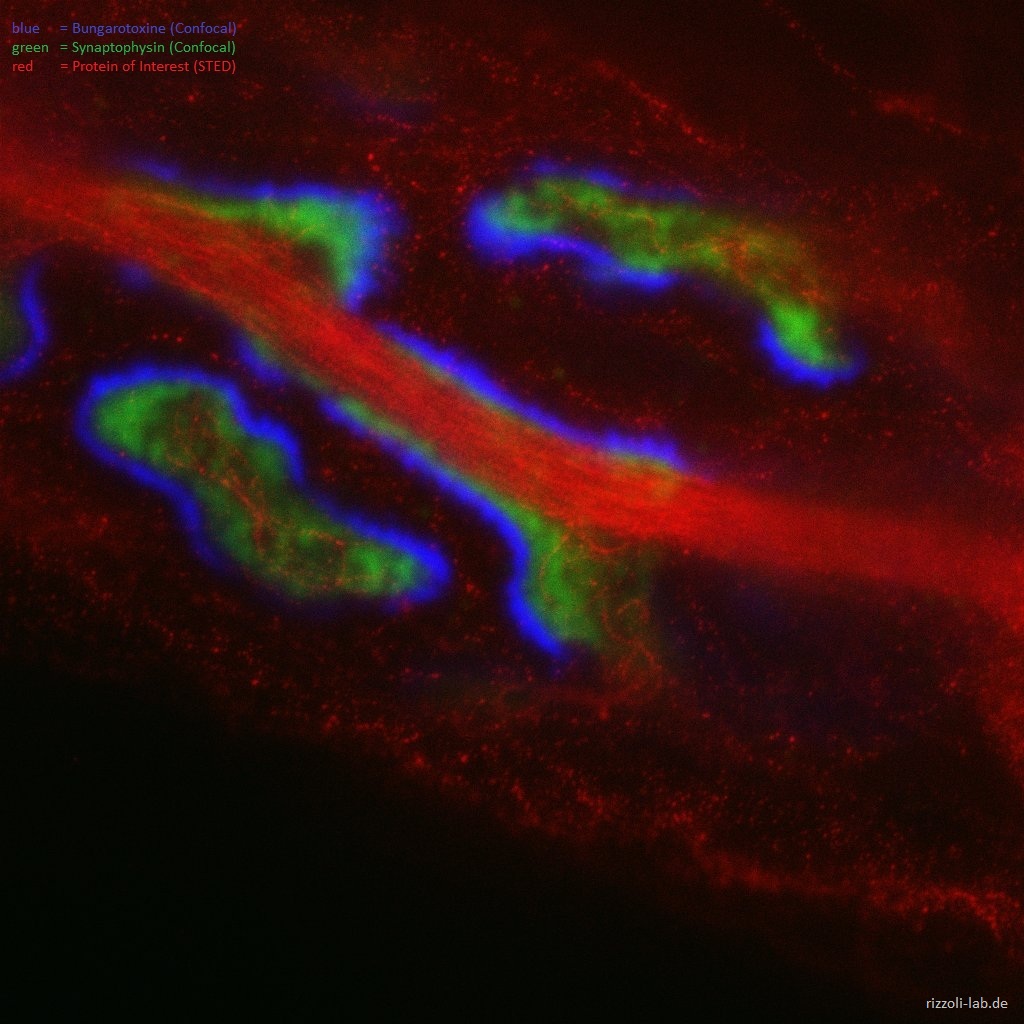
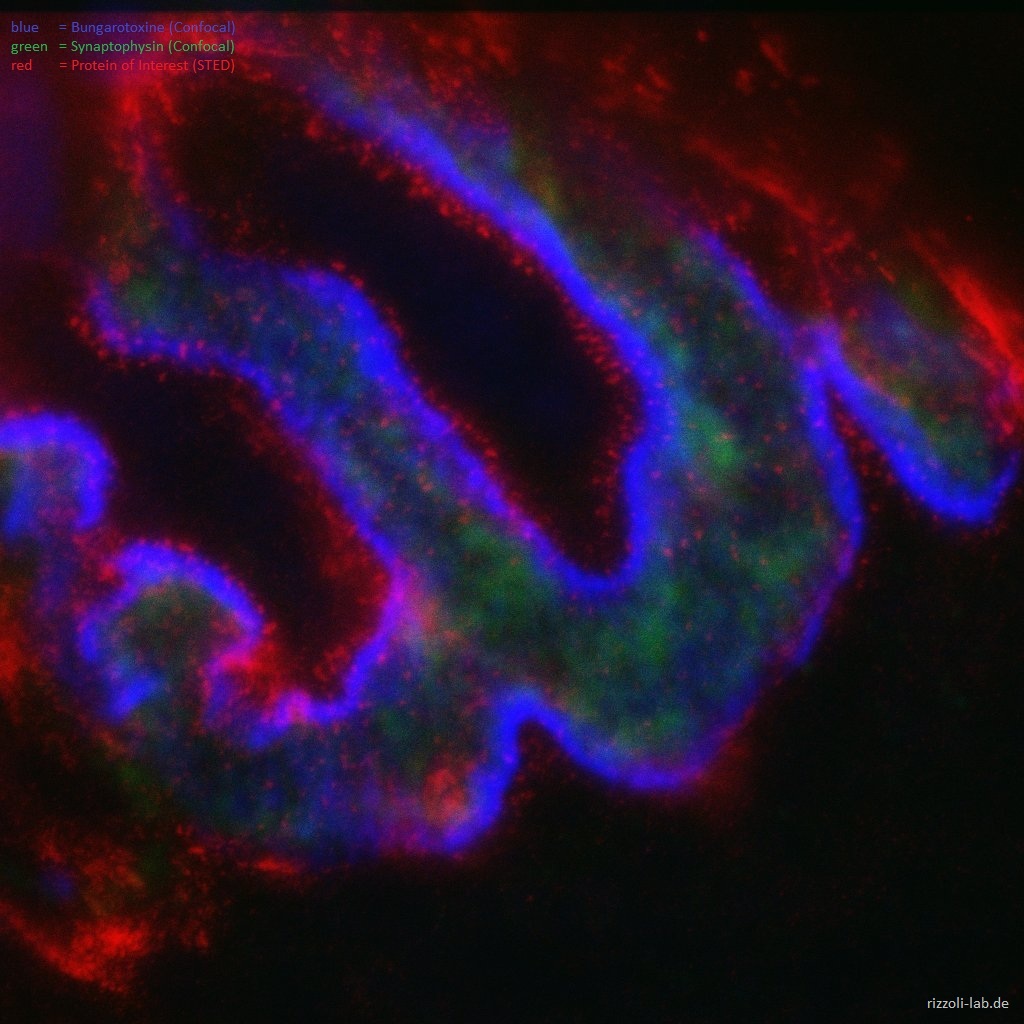
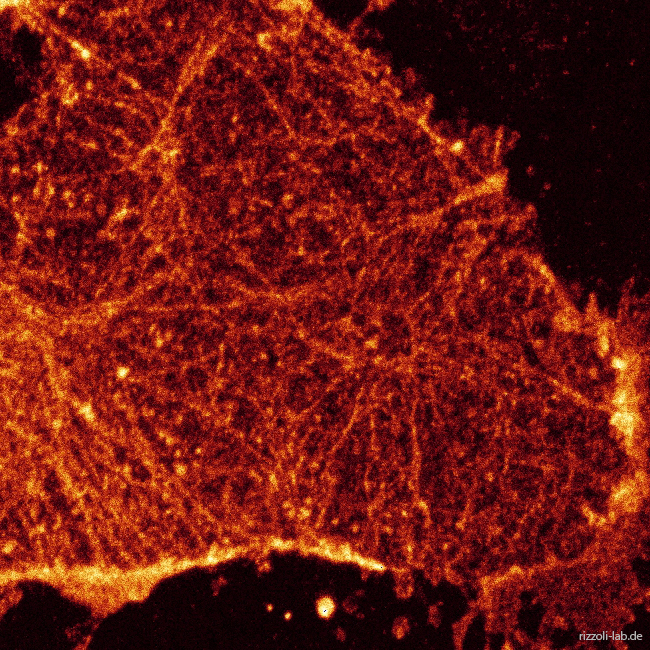
Our laboratory has a dual focus – high-end imaging and cutting edge research in synaptic physiology. All projects make use of advanced imaging techniques, including STED, electron microscopy and NanoSIMS. The ultimate goal of our work is to understand the functional organization of the cell – the connection between the topological distribution of cellular elements (proteins or organelles) and their function. We currently focus on the presynaptic compartment, whose relative simplicity and well understood function render such studies more feasible.
Briefly, most proteins investigated so far by high resolution imaging form super-molecular assemblies or clusters. The reasons why they do so are complex, with no general explanation available, especially as the location of the clusters often does not correlate with the protein function.
For example, fusion proteins involved in synaptic vesicle exocytosis can be found in clusters everywhere on the plasma membrane, even in places where there are no vesicles. Having an excess number of copies in a cluster is not limited to proteins: synaptic vesicles can be found clustered in immense numbers, especially in neuromuscular junctions, with few explanations available as to the functional relevance of most vesicles (for example, from the ~500,000 vesicles in the frog cutaneous pectoris neuromuscular junction, only ~250 vesicles are used for one action potential).
Thus, the functional organization of many synaptic (or cellular) elements is poorly understood. To solve this type of question, we combine imaging, biochemistry and biophysics to determine the amount of free molecules, of molecules found in clusters, as well as their exchange rates. We then compare all of these values to the numbers of molecules used in function. We termed this approach stoichiometric biology (Lang and Rizzoli, 2010), and we apply it to study both protein clusters and organelle (vesicle) organization. We have recently published the first demonstration of this approach, proposing a novel function for the clusters of synaptic vesicles, in buffering soluble synaptic proteins (Denker et al., 2011b).

Imageomics Future and Emerging Technologies (FET-OPEN) 2020: High-throughput imaging of proteins using nanobodies Horizon 2020 01.07.2021-31.12.2024 The conventional detection of proteins by imaging employs fluorophores
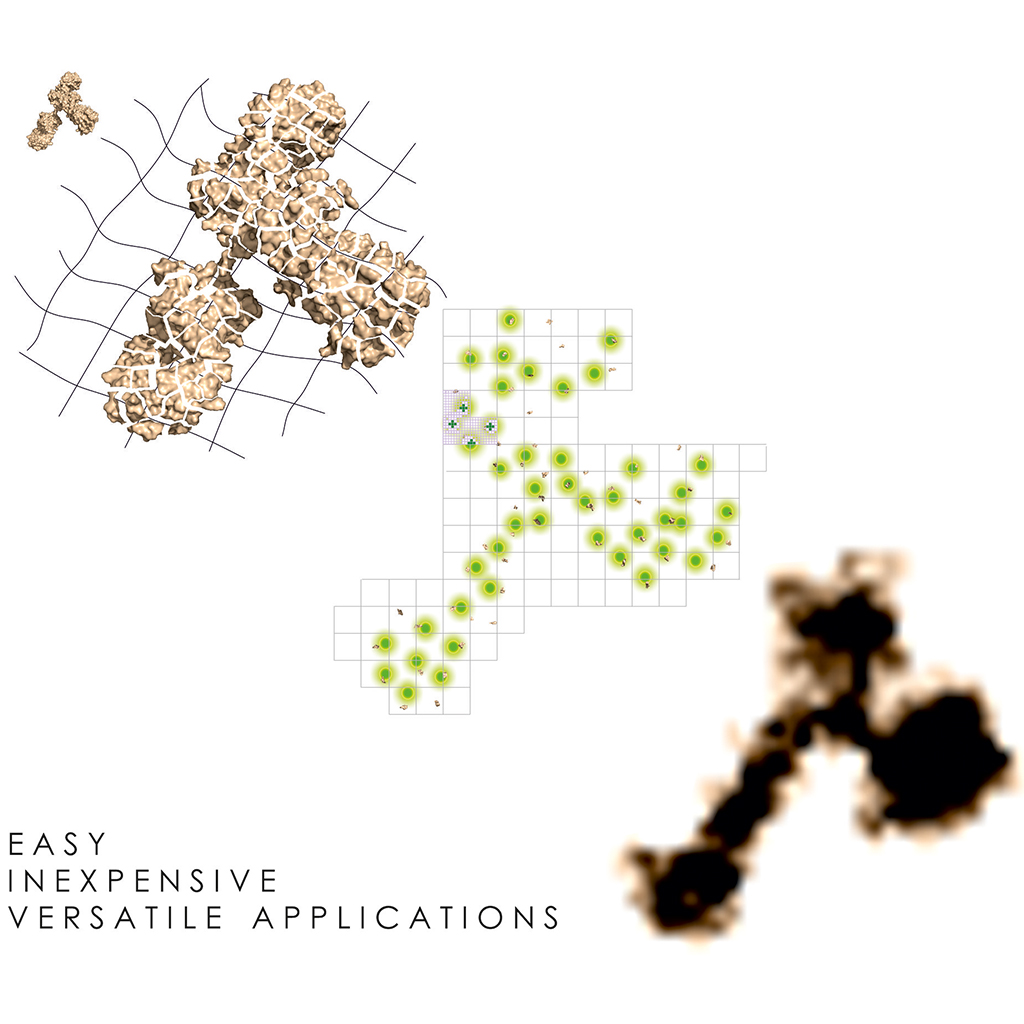
ONE Microscopy Optimized Nanoscale Expansion microscopy Super-resolution imaging has been applied to biological specimens for more than two decades. Nevertheless, it still falls short of
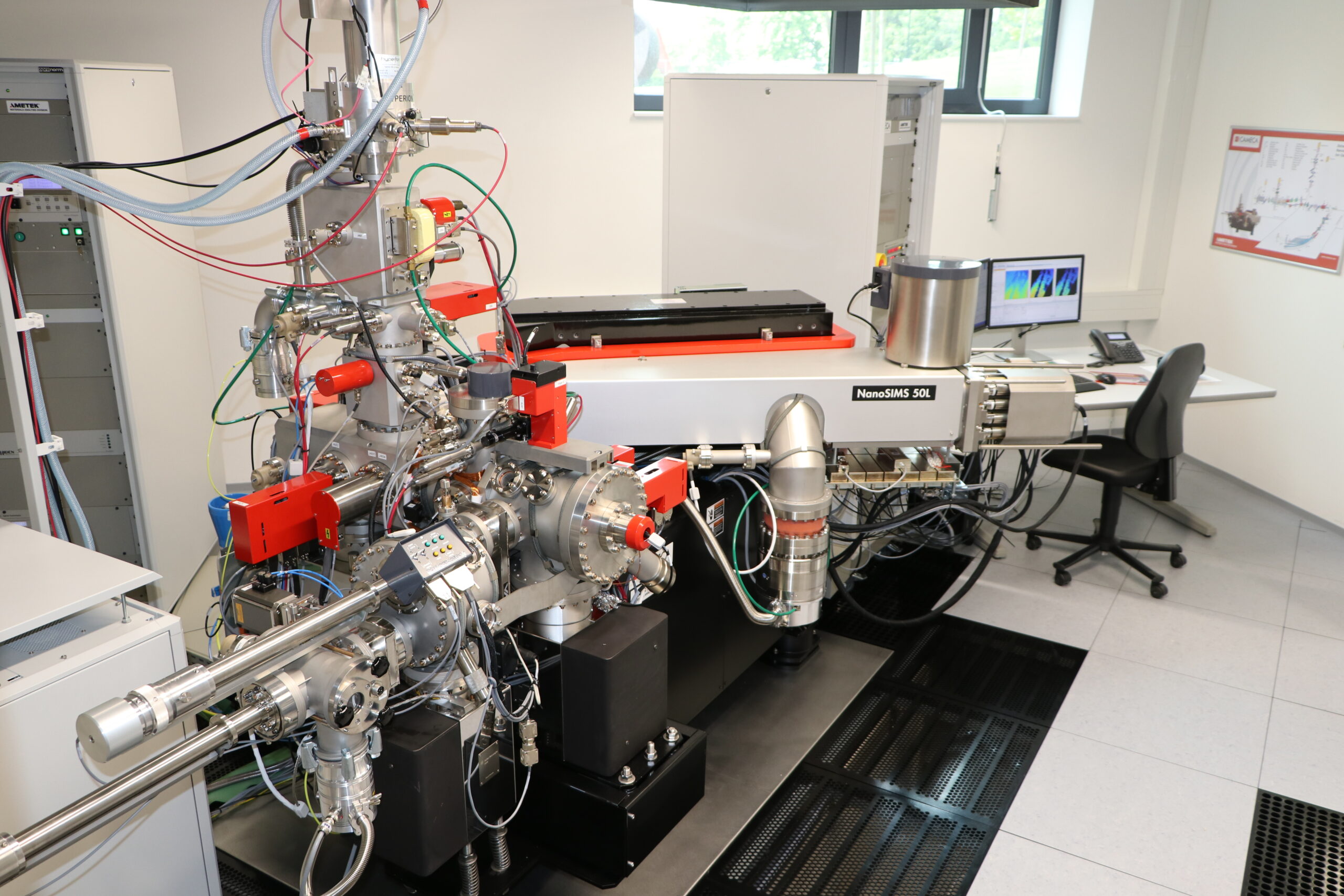

NanoSIMS 50L Starting at the end of 2017, the Center for Biostructural Imaging of Neurodegeneration (BIN) can offer the services of a new high performance

ULTRARESOLUTION Beyond super-resolution: ultra-resolution imaging provides solutions for synapse physiology and brain pathology ERC Synergy Grant 2020 01.06.2021-31.05.2027 Neurons contain hundreds of specialized proteins, whose
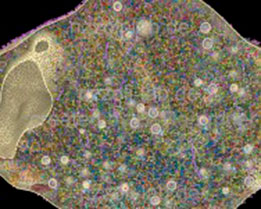
The Neuron Nanomap In this project, which has started in April 2014, we generate a super-resolution image – a nanomap – of presynaptic nerve terminals,